Lab 3: Two-Predictor Regression
PSYC 7804 - Regression with Lab
Spring 2026
Today’s Packages and Data 🤗
The GGally package (Schloerke et al., 2024) builds upon ggplot2 and includes many functions for creating complex plots.
The devtools package (Wickham et al., 2022) helps developing R packages. It is generally used to download R packages from github.
This is my personal package where I save functions that I use for research/teaching/convenience. You can run devtools::install_github("quinix45/FabioFun") to install it.
The flextable package (Gohel et al., 2024) allows to create highly customizable tables. The function that I use to create APA style Word tables makes heavy use of this package.
Predicting Country Happiness
In our data we have many variables that are likely related to happiness. I will just pick 2 of them for the purpose of this Lab. Let’s say that we want to evaluate how Freedom and Corruption predict Happiness. If we run two separate regressions, we find that both have reasonably high \(R^2\) values:
Call:
lm(formula = Happiness_score ~ Freedom, data = dat)
Residuals:
Min 1Q Median 3Q Max
-2.5273 -0.5317 0.1337 0.6656 1.7318
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.6234 0.3035 8.644 1.22e-14 ***
Freedom 4.6849 0.4732 9.901 < 2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.9065 on 138 degrees of freedom
Multiple R-squared: 0.4153, Adjusted R-squared: 0.4111
F-statistic: 98.03 on 1 and 138 DF, p-value: < 2.2e-16
Call:
lm(formula = Happiness_score ~ Corruption, data = dat)
Residuals:
Min 1Q Median 3Q Max
-3.5303 -0.7029 0.3538 0.7954 1.8762
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 9.1072 0.6077 14.99 < 2e-16 ***
Corruption -4.2279 0.7106 -5.95 2.1e-08 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.058 on 138 degrees of freedom
Multiple R-squared: 0.2041, Adjusted R-squared: 0.1984
F-statistic: 35.4 on 1 and 138 DF, p-value: 2.1e-08Well, That’s just the correlation 🤷
As we have learned, regression with one predictor is just a contrived way of calculating correlations among variables. We can better visualize the relation among out 3 variables with the ggpairs() function from GGally:
Separate regressions miss the points that our predictors share information!
Multiple Independent Variables
Call:
lm(formula = Happiness_score ~ Freedom + Corruption, data = dat)
Residuals:
Min 1Q Median 3Q Max
-2.3574 -0.4326 0.1503 0.5999 1.7335
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 5.0938 0.6875 7.409 1.18e-11 ***
Freedom 4.0320 0.4792 8.415 4.63e-14 ***
Corruption -2.4415 0.6168 -3.959 0.000121 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.8618 on 137 degrees of freedom
Multiple R-squared: 0.4753, Adjusted R-squared: 0.4677
F-statistic: 62.06 on 2 and 137 DF, p-value: < 2.2e-16We can include both predictors in our regression by adding variables in the lm() function like so lm(DV ~ IV1 + IV2 + ..., dat).
Note that both slopes have changed and are not as large as they were in the individual regressions. The regression equations is now:
\[\mathrm{Happiness} = 5.09 + 4.03 \times \mathrm{Freedom} - 2.44 \times \mathrm{Corruption}\]
What is going on here? 🤔 a graphical illustration may provide some intuition!
Individual regression plots
Below I plot the individual regression lines for Freedom and Corruption predicting Happiness.
Now, what happens if we try to overlay the regression lines that we got from our multiple regression? Will they “fit the data” better that the regression lines shown here?
Adding Multiple regression Lines
if we add our multiple regression lines to the plot… (🥁 drum roll)
Wait, we are way off now 🤨 And here I thought multiple regression would be better 😑 …Unsurprisingly, the multiple regression is fine, our perspective is the problem!
A better Perspective
We have 3 variables, Freedom, Corruption, and Happiness, but we are looking at them in 2 dimensions with the individual scatterplots. We need a 3D scatterplot!
What is happening is that the scatterplots are only the sides of this box ‼️
Multiple regression works on this plot on the right, not the individual ones; multiple regression operates on a different dimension.
The Regression Plane
The results from multiple regression create a regression plane, which is the 3D version of a regression line.
The plane works exactly as the regression line. its surface is the best predition of happiness given a value of Freedom and Corruption. The residuals are now the distance from any of the dots to the plane!
Question: What would the plot look like with one extra variable?
Interpreting Multiple Regression
Call:
lm(formula = Happiness_score ~ Freedom + Corruption, data = dat)
Residuals:
Min 1Q Median 3Q Max
-2.3574 -0.4326 0.1503 0.5999 1.7335
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 5.0938 0.6875 7.409 1.18e-11 ***
Freedom 4.0320 0.4792 8.415 4.63e-14 ***
Corruption -2.4415 0.6168 -3.959 0.000121 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.8618 on 137 degrees of freedom
Multiple R-squared: 0.4753, Adjusted R-squared: 0.4677
F-statistic: 62.06 on 2 and 137 DF, p-value: < 2.2e-16Now that we have looked at our results graphically for a while, let’s interpret the numbers that we get.
intercept: the expected value of
Happinesswhen bothFreedomandCorruptionare at 0.Slope of
Freedom: the expected change inHappinessfor 1-unit increase inFreedom, controlling forCorruption.Slope of
corruption: the expected change inHappinessfor 1-unit increase inCorruption, controlling forFreedom.
Note: Instead of “controlling for”, you may also hear “accounting for” or “holding constant”. They are all the same thing.
Variance as what we cannot predict
At this point, a quick detour about variance is in order. In Psychology we work with random variables. The word random is important because it means that there is something that we cannot predict about the variable (e.g., we cannot perfectly predict depression, nor will we ever be able to).
However, given a sample, we can quantify the unpredictability of a variable. The variance (or equivalently the SD) measure how uncertain a variable is!
We can see that the red distribution has a larger variance than the blue distribution. That is, the red distributrion has more unpredictability.
(by the way, generally Entropy is used to quantify unpredictability of a random variable)
Plot code
ggplot() +
geom_function(fun = dnorm,
args = list(mean = -4,
sd = 1), color = "blue") +
geom_function(fun = dnorm,
args = list(mean = 4,
sd = 2), color = "red") +
theme(axis.title.y=element_blank(),
axis.text.y=element_blank(),
axis.ticks.y=element_blank(),
axis.line.y = element_blank()) + xlim(c(-10, 10))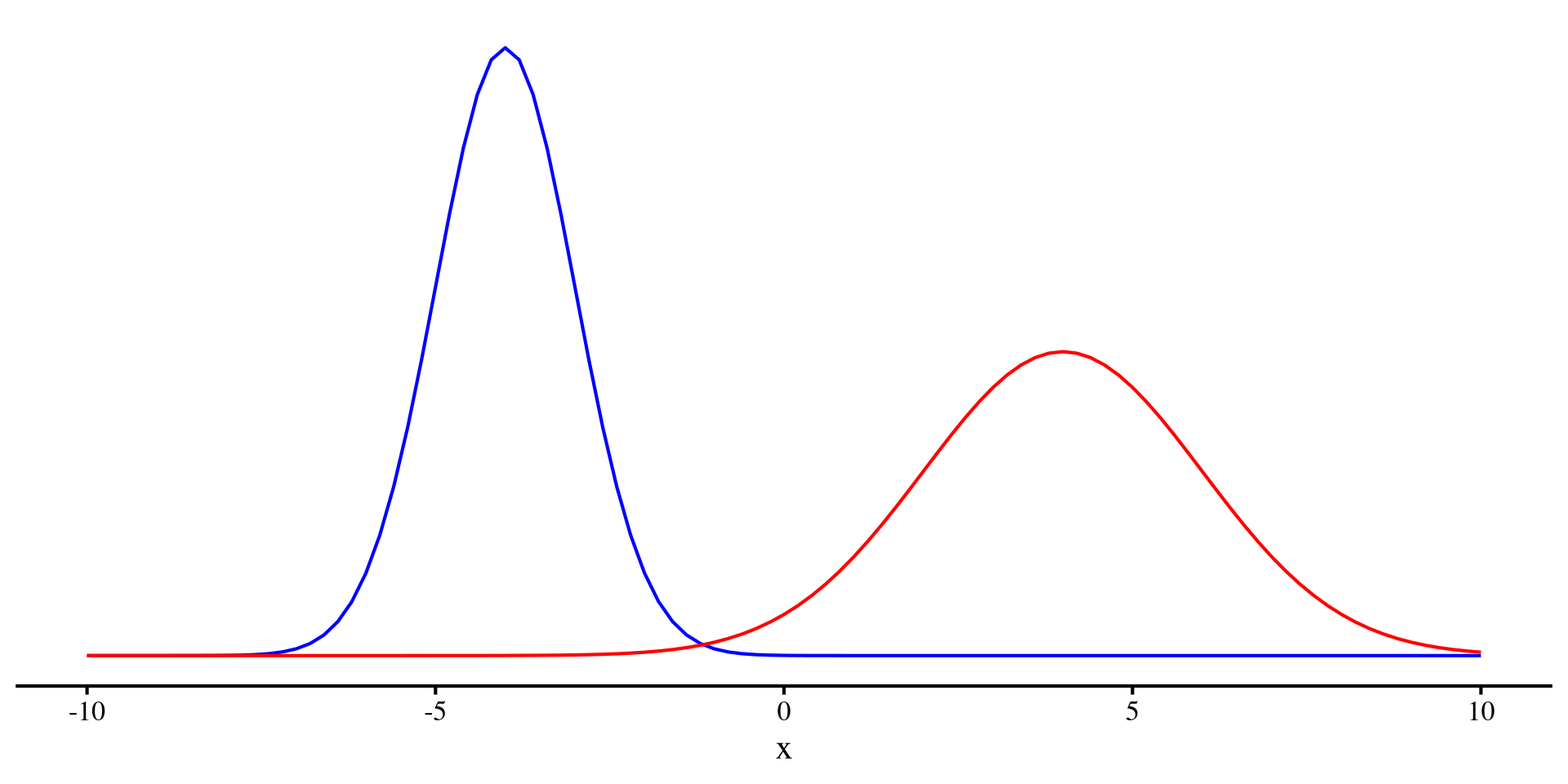
R2 as Uncertainty Explained
After running a regression we can calculate a statistic called \(R^2\), which quantifies the predictive power of the regression. \(R^2\) is also often termed “variance explained”. Let’s figure out why:
Our regression where \(Y =\) Happiness had an \(R^2\) of:
So, we can see that the variance of our \(Y\) variable, Happiness, has been reduced by
\[ 1 - \frac{0.73}{1.395} = 0.475\% = R^2 \]
\(R^2\) is an effect size and it tells us how much uncertainty in the \(Y\) variable has been reduced by the predictors.
A graphical look at R2
We can plot the density of Happiness and the residuals just like we plotted the two distributions a few slides ago.
Just as before, you can see that the two distributions visibly differ in variance. The residuals (blue density) have less variance, \(47.5\%\) less to be precise.
Question: What would it mean if the residuals had no variance at all? 🤔
Plot code
ggplot(dat) +
geom_density(
aes(x = Happiness_score, color = "Happiness (Original Variable)")
) +
geom_density(
aes(x = reg_multi$resid, color = "Regression residuals")
) +
scale_color_manual(
name = "",
values = c(
"Happiness (Original Variable)" = "red",
"Regression residuals" = "blue"
)
) +
theme(
axis.title.y = element_blank(),
axis.text.y = element_blank(),
axis.ticks.y = element_blank(),
axis.line.y = element_blank(),
legend.position = "bottom"
) +
xlim(c(-5, 10))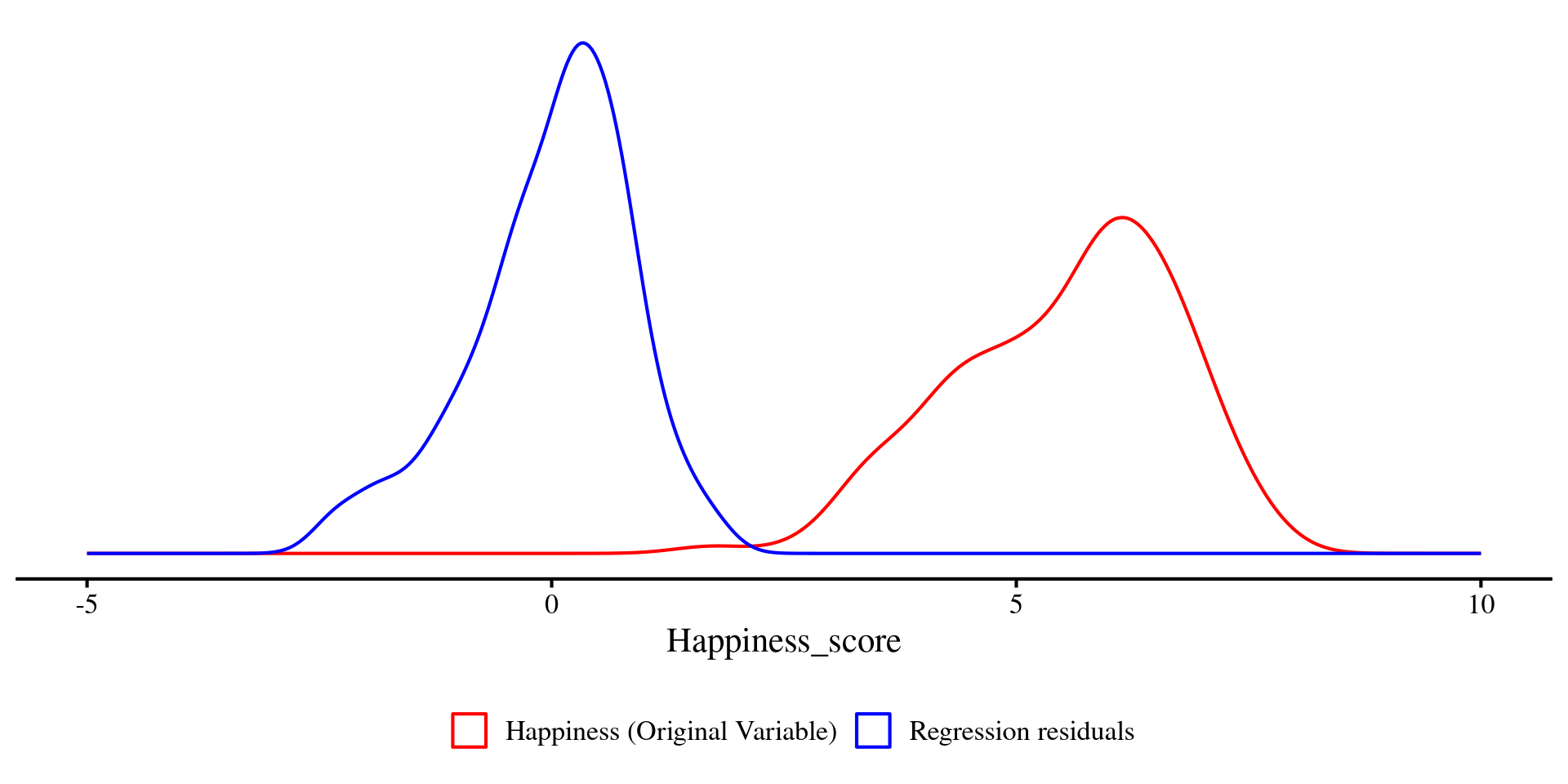
Significance Test for R2
We have seen that we can individually test whether each of the regression coefficients are significantly different from 0 with t-tests. However, this may become a bit cumbersome when we introduce many predictors in our regression.
If you are not as interested to every individual variable, another approach is to test whether the regression \(R^2\) is significantly different from 0.
The significance test for the regression \(R^2\) is also found at the end of the summary() output. Here we have that \(R^2 = .48, F(2, 137) = 62.06, p < .001\).
This means that our regression model explains a statistically significant proportion of variance in the outcome, Happiness.
APA Style Table for Regression Results
because I hate creating tables manually, at some point I made function that does that for me ✨ Also, creating big tables by hand is generally not a good idea: you waste time and likely make typos.
the regression_flextable() function I made takes in an lm() object and has some optional arguments (see help page). Here I rename the regression coefficients with the var_names = argument.
You can save the table as a .docx file (Word document) by using the save_as_docx() function from the flextable() package. You specify the file name in the path = argument.
References
Appendix: nice_3D_plot() Showcase
Example with World Happiness Data
You can also add groups to the dots of the 3D plot. Here I am using Region.
# selecting just the variables of interest makes code cleaner later
reg_vars <- dat[, c("Happiness_score", "Log_GDP", "Freedom")]
nice_3D_plot(y = reg_vars$Happiness_score,
x1 = reg_vars$Log_GDP,
x2 = reg_vars$Freedom,
dot_labels = dat$Country_name,
# add groups
groups = dat$Region,
axis_names = c("Happiness",
"GDP",
"Freedom"),
plane_res = 20,
reg_plane = FALSE)There are a bit too many regions, so this is not very practical in this case, but this just an example.
Example with iris Data
A more practical example with the iris dataset, Let’s start with just providing 3 variables.
Name Axes
Let’s name our axes:
Add Groups
Let’s add colors for the 3 different species of flowers:
Add Regression Plane
Let’s add a regression plane:
Pass Arguments to plot_ly()
You can also pass arguments to the plot_ly() function. Let’s make the dots more transparent by passing the opacity = argument to plot_ly():
See here for what “passing arguments” to a function within a function means in R.
Add Interaction Term to Regression Plane
You can also specify an interaction between the two predictors and get an interaction plane. You will see that the reression plane now bends:
See Lab 9 if you are curious to know what is happening.
PSYC 7804 - Lab 3: Two-Predictor Regression




